

Ongoing Projects.

Domain Specific Many-Core
Team Members: Aakarsh Vinay, Hardhik Benedict, Amith V, Suravarjhula Sindhura, Ashuthosh M.R
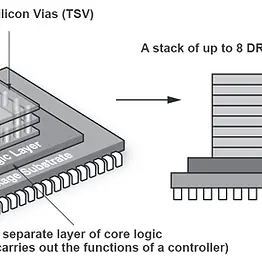
In-Memory Computation (IMC) using Hybrid Memory Cube (HMC)
Team Members: Gowtham D, Shalett Paulson, Mohan Krishna N D, Lokesh and Chandan

Instruction Profiler - RISC-V Profiling for identification of custom instruction-set extensions
Team Members: Nandhini Dhanasekharan, Nikhita Sridhar, Shruti Narayana

PSIMD Extension of RISC-V
Team Members: Dinesh Venkat G, K S Vandana, Adrika Mohanty, R Mauriya, Vivek R, Pooja Tirmal, Rishab Somani

Physical Hardening of RISC V
Team Members: Vinayaka M Kanti, Prajna S, Prathiksha

RAP: RISC-V Application Profiler
Team Members: Mahendra Vamshi, Sowmya Ravi, MD Sabir

Sub-Threshold Standard Cell Design
Team Members: Vardhan, Yagna Vivek, Sashank Sharma, Ishitha Aagarwal, Vikram Kannur
Past Projects.

Accelerating Molecular Dynamics
Within this project, our objective is to identify the drawbacks in current FPGA architecture, since they have been mainly targeted for DSP applications and algorithms. Some questions that we want to answer are (1) What are the features that FPGAs lack that could simplify/accelerate drug discovery 1 applications? (2) How can FPGAs help accelerate molecular dynamics (3) Could partial reconfiguration be a boon for computational chemists? (4) Could we re-architect the FPGA specifically for drug discovery applications?

Circuit-level exploration of FPGA architectures
Transistor sizing in FPGAs is a complex optimisation problem. Studies in this direction have explored the impact of sizing closely coupled Lookup tables (LUT) in identical tile-based FPGAs. Our focus is to analyse the impact of application-level variabilities introduced through the configuration data or input changes. In this paper, our objective is to: (1) understand the impact of application-level data on transistor sizing in pass-transistor-based LUTs and (2) suggest an alternative LUT implementation that guarantees constant response time.

Hardware accelerators for Deep Learning
Convolution Neural Networks (CNNs) are becoming increasingly popular in Advanced driver assistance systems (ADAS) and Automated driving (AD) for camera perception enabling multiple applications like object detection, lane detection and semantic segmentation. Ever increasing need for high resolution multiple cameras around car necessitates a huge-throughput in
the order of about few 10’s of TeraMACs per second (TMACS) along with high accuracy of detection. This project will suggest an architecture that is scalable exceeding few 100s of GOPs.

Accelerating Genome Sequence Analysis
In computational genomics, the term kmer typically refers to all the possible subsequences of length k from a single read obtained through DNA sequencing. In genome assembly, generating frequency of k-mers takes the highest compute time. k-mer counting is considered as one of the important analyses and the first step in sequencing experiments. Here, we explore an FPGA based fast k-mer generator and counter,k-core to generate unique k-mers and count their frequency of occurrence.

Graph Algorithms
We considered a few popular graph algorithms– PageRank, Single Source Shortest Path (SSSP), Breadth-First Search (BFS), and Depth-First Search (DFS). We employed the High-level synthesis and its optimization methodologies to design an FPGA accelerator for the respective algorithms. Due to resource constraints on the device, we adopted algorithm-specific graph partitioning schemes to process large graphs. Using the GAP Benchmark Suite running on a CPU as the baseline for evaluating the performance of our design we obtained a speedup of 5x for BFS, 20x for SSSP.
Source code and reports are here

Accearator for Genomics
Genome sequencing is increasingly used in healthcare and research to identify genetic variations. The genome of an organism consists of a few million to billions of base pairs. Oxford Nanopore sequencing works by monitoring changes in electrical current and the resulting signal is basecalled using a neural network to produce the DNA sequence. Deep Learning operations are computationally intensive because they involve multiplying tensors (multi-dimensional matrices). We use methods like pruning and quantization to lower the amount of computation and reduce the size of the model.

Cardiac Anomaly Detection
Sudden Cardiac Arrest (SCA) is a devastating heart abnormality which leads to millions of casualties per year. Thus, early detection or prediction of SCA could save human lives in a greater scale.
In this work, we aim to predict SCA before its occurrence and significant results has been obtained using the proposed signal processing methodology. Models were trained using a CNN, CNN + Long Short Term Memory (LSTM) model and a Random Forest Classifier.
Team Members: Anktih B V, Aaptha B V and Aryan Sharma

